A Detection of Informal Abbreviations from Free Text Medical Notes Using Deep Learning
2 Graduate School of Medicine, Kyoto University, Japan, Email: kuhp.kyoto-u@ac.jp
3 Kyoto University Hospital, Japan, Email: kuhp.kyoto-u@ac.jp
4 Research Center for Informatics, Indonesian Institute of Sciences, Indonesia, Email: lipi@go.id
Citation: Heryawan L, et. al. (2020). A Detection of Informal Abbreviations from Free Text Medical Notes Using Deep Learning. EJBI. 16(1): 29-37
This open-access article is distributed under the terms of the Creative Commons Attribution Non-Commercial License (CC BY-NC) (http://creativecommons.org/licenses/by-nc/4.0/), which permits reuse, distribution and reproduction of the article, provided that the original work is properly cited and the reuse is restricted to noncommercial purposes. For commercial reuse, contact submissions@ejbi.org
Abstract
Background: To parse free text medical notes into structured data such as disease names, drugs, procedures, and other important medical information first, it is necessary to detect medical entities. It is important for an Electronic Medical Record (EMR) to have structured data with semantic interoperability to serve as a seamless communication platform whenever a patient migrates from one physician to another. However, in free text notes, medical entities are often expressed using informal abbreviations. An informal abbreviation is a non-standard or undetermined abbreviation, made in diverse writing styles, which may burden the semantic interoperability between EMR systems. Therefore, a detection of informal abbreviations is required to tackle this issue.
Objectives: We attempt to achieve highly reliable detection of informal abbreviations made in diverse writing styles.
Methods: In this study, we apply the Long Short-Term Memory (LSTM) model to detect informal abbreviations in free text medical notes. Additionally, we use sliding windows to tackle the limited data issue and sample generator for the imbalance class issue, while introducing additional pre-trained features (bag of words and word2vec vectors) to the model.
Results: The LSTM model was able to detect informal abbreviations with precision of 93.6%, recall of 57.6%, and F1-score of 68.9%. Conclusion: Our method was able to recognize informal abbreviations using small data set with high precision. The detection can be used to recognize informal abbreviations in real-time while the physician is typing it and raise appropriate indicators for the informal abbreviation meaning confirmation, thus increase the semantic interoperability.
Keywords
Informal abbreviations; LSTM; Structured data; Free text medical notes; EMR
Introduction
Medical data need to be structured to achieve semantic interoperability. Semantic interoperability is essential for Electronic Medical Records (EMR) since they must serve as a seamless communication platform, allowing data to be compatible whenever a patient migrates from one physician to another [1,2]. Semantic interoperability ensures that the meaning of medical concepts can be shared across systems, thus providing a digital and common language for medical terms that is understandable to humans and machines.
For instance, the sentence “patient g2-p2 experiences asthma attack” includes information about pregnancy history, in the form of the abbreviated term “g2-p2”. If the same sentence were written using a standard for semantic interoperability, such as SNOMED CT [3], the abbreviation “g2-p2” would be replaced by “gravida 2 or second pregnancy and para 2 or parity 2”. In the abbreviated sentence, if the term “g2-p2” were not detected by a healthcare information system that employs SNOMED CT, such as a Clinical Physician Order Entry (CPOE), the person responsible for processing the medication order might misunderstand or fail to recognize it, leading to an erroneous interpretation and inappropriate drug administration, which increases the risk for the patient.
Additionally, a large amount of medical data produced currently are free text medical notes [4], such as SOAP notes and discharge summaries. These documents are often created in varied writing styles [5], using informal abbreviations that do not follow standard medical nomenclature [6]. Since these documents are implicitly composed in an unstructured format, it is necessary to parse them and identify specific information in the text to finally generated structured data that can be shared among different systems. In this process, detecting abbreviations correctly is a key requirement since terms unknown to the system could be ignored or misrepresented in the final output.
The problem of detecting abbreviations falls in one of natural language processing tasks, which is named entity recognition (NER). NER task is identifying specific terms designating medical concepts within the text [7], in our case is identifying the presence of abbreviations. This is done in two steps: abbreviation detection and then abbreviation disambiguation, which is a special case of word sense disambiguation (WSD) [8]. In this study, we focus on the first step, abbreviation detection, which remains a challenging problem, due to the variability of the input.
A previous study [9] showed the potential usefulness of machinelearning- based (ML-based) abbreviation detection methods for predefined abbreviations in English texts and medical notes. Hence, that study focused on detecting formal abbreviation with predefined extracted features using traditional machine learning algorithms, such as decision trees, random forests, and support vector machines. Another previous study to detect abbreviation also used machine learning algorithm, such as stochastic gradient descent [10]. While deep learning algorithm such as bidirectional LSTM and CNN are used to detect more complex named entity such as nested named entity [11] and biomedical named entity [12]. In contrast, our study focuses on detecting informal abbreviations that is a noisy entity, which has not been covered by the previous study.
To detect such informal abbreviations, we propose a Long Short-Term Memory (LSTM) based model [13], a form of deep learning technique. Deep learning algorithms a subset of machine learning methods-currently offer the best performance in tasks that involve learning from sequential data, such as free text medical notes. There are several deep learning algorithms for the information extraction task, such as Convolutional Neural Network (CNN), Recurrent Neural Network (RNN), Recursive Neural Network, Neural Language Model, and Deep transformer [14]. The information that can be extracted from EMR using deep learning are varied, from a patient phenotyping using CNN [15] to named entities using RNN [16]. In our informal abbreviation extraction study, we choose RNN variant, an LSTM based model as our proposed model. It learns without the need of prior features extraction processes [17]. Although the feature extraction task is particularly difficult in the case of abbreviations detection, the LSTM model can be effective, since it also takes into consideration the context in which sequences occur to detect the intended entity at each location. In other words, the model can detect abbreviations by exploring their surroundings. For example, the LSTM model is able to detect that “G3P3” is an abbreviation in the sentence “pregnancy-labor-spontaneous delivery to a G3P3”, as is the case of “g3.p2A1” in the sentence “pregnancy-labor-spontaneous delivery to a g3.p2A1”. It does so by understanding that these words occur in similar contexts.
This study presents deep learning modelling using LSTM to detect informal abbreviations from medical notes sentences, which is a vital task for semantic interoperability in EMR. Additionally, our approach expands the field of application of the LSTM model since the technique was not used before for this kind of task.
Methods
In order to detect the informal abbreviations in the free text medical notes, we propose an LSTM based method. The proposed method has components corresponding to the following problems, first as a detection of noisy entities, i.e., terms to be identified in a set of unstructured data. Second, the detection task suffer from the problem of class imbalance, in which different types of entities have very difference frequency of occurrence in the data, making some of them underrepresented.
Free text medical notes are an example of this kind of data. A similar case is that of Twitter messages (tweets [18]). Both types of data consist of noisy entities that lack implicit linguistic formalism (e.g. improper punctuation, spelling, spacing, formatting) while also including abbreviations [19]. The LSTM model outperforms other strategies when used for tasks of entity recognition in this kind of noisy data set.
Noisy entities are particularly difficult to detect if the surrounding context is not considered, including the order and relative distribution of the entity’s occurrence. The entity’s order can be represented by a word2vec matrix, while the entity’s relative distribution can be represented by Bag of Words (BoW) matrix. For this reason, the addition of these representations to the input of the model can increase its accuracy.
Additionally, the problem of class imbalance must be addressed. It happens when the total number of items in a data class (in this case, abbreviations) is far less than the total number of items in another class (non-abbreviation). To deal with this problem, one common approach is to use oversampling [20], which consists of increasing the number items of the under-represented class, so it has a greater impact on the machine learning algorithm.
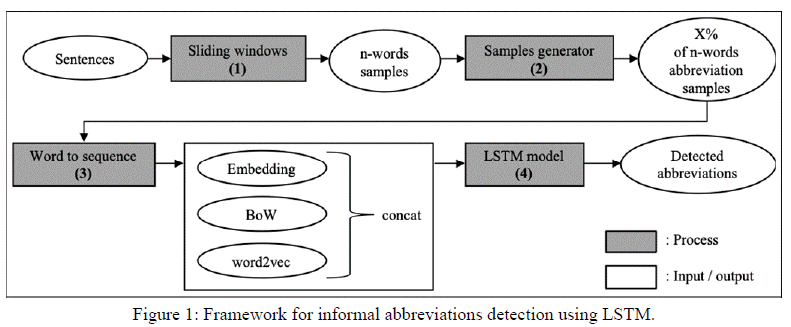
Our method consists of two parts: pre-processing, with data oversampling to address class imbalance; and processing, with additional inputs to represent word context (word2vec, BoW). Figure 1 illustrates the detection flow using our framework.
The pre-processing part is further divided into two steps, sliding window and samples generator. The first step (sliding window) involves breaking the longer and varied-length sentences into fixed groups of words. This is necessary because the LSTM model requires short fixed length sequential inputs. This transformation also alleviates the problem of limited data sets (Section 5.4), since it increases the number of samples. The second phase of preprocessing (samples generator) increases the number of samples of infrequent data [21], to improve the LSTM model’s prediction performance, since it performs better with more balanced data.
The strength of LSTM model is that it learns its word embedding input automatically, although it can also learn from a pre-trained word embedding. A pre-trained word embedding is represented as a matrix, which can be produced using a word representation algorithm, such as BoW and word2vec. In our abbreviation detection, we concatenate word embedding with the pretrained embedding, BoW matrix and word2vec matrix (step 3) to achieve higher performance. This takes advantage of the fact that the LSTM model (step 4) can process multiple features for its learning [22].
Pre-processing
The pre-processing process was designed for solving the problems of class imbalance and unfair learning of the LSTM classifier. The issue causes a low predictive performance of classifier to recognize the minority class.
To tackle the class imbalance issue, we use a sliding window [23] to split the input sentence into chunks, and a samples generator to increase the number of abbreviation samples. The sliding windows step creates fixed-length samples with size n, consisting of a sequence of words from the sentence, without any changes in the words themselves. In our case, we use window size 5 (n=5), and step 1 (the next window starts 1 word to the right), the input sentence “coronary artery disease history of seizure disorder GERD bipolar” would be chunked into five fixed size samples, starting with “coronary artery disease history of”, then “artery disease history of seizure”, and so on. We manually labelled words as abbreviation/non- abbreviation for the training process.
After fixed size inputs are produced, a samples generator is used to increase the ratio of abbreviation samples in the data set. This step is necessary to create fairer learning for the LSTM model, by reducing class imbalance from the input. By specifying the percentage of samples that should contain at least one abbreviation word, we determine the ratio of abbreviation/ non-abbreviation samples present in the input presented to the model.
Processing
The processing stage was designed with the consideration of sentence feature extraction. In this stage, we use a LSTM with one embedding layer, which is used as our baseline model. The common approach when using the LSTM model for entity classification from text is to convert each word in the sample to a word index, which is a positive integer. After that, each word index is turned into embedding vectors of fixed size [24] before being presented as input to the LSTM model.
A bidirectional LSTMs algorithm is used for our detection of informal abbreviations. The bidirectional LSTMs are an extension of traditional LSTMs that can improve model performance on sequence classification problems [25]. In problems where all time steps of the input sequence are available, such as free text sentences, the bidirectional LSTMs train two instead of one LSTMs on the input sequence. The first on the input sequence asis and the second on a reversed copy of the input sequence. This can provide additional context to the model and result in faster and even fuller learning on the problem.
We trained the model using additional pre-trained matrices such as word frequency or BoW matrix [26], and word2vec matrix [27] to increase our model performance. The BoW matrix represents the occurrence frequency of each word in relation to the complete vocabulary in the data set. The word2vec matrix represents words as vectors in such a way that words which share similar contexts are closer in the vector space [28]. For example the BoW matrix of the word “5” in the sentence “A 5 yr boy brought by his parents because of 2 days of cough” is (0,1,0,0,0,0,0,0,0,0,0,0,0,0) and the vector for the word “5” is clustered together with the vector for the word “2”.
Based on the fact that the LSTM model can process multiple features to increase its performance [22], the additional matrices are concatenated with the embedding vectors of the baseline model. The BoW and word2vec matrices were used as additional input to the LSTM model based on the idea that abbreviation words have frequencies and vectors in the word2vec vector space that differentiate them from non-abbreviation words and ordinary English words. It is expected that the model’s performance would be increased with the inclusion of more relevant pre-trained features, represented by these additional matrices.
In summary, our basic LSTM model used only embedding vectors as an input. The other input combinations we evaluated expanded this basic model by including additional preTrained features in the form of the BoW and word2vec matrices. To enable this additional input to the LSTM model it was necessary to include additional layers for each of the additional matrix of features [29]. Finally, all the LSTM layers in the model are concatenated to predict the abbreviation in the sample using LSTM hyper- parameters, such as dropout, dense, and activation layer. In our model, the hyper parameters are the same for all inputs combination.
Experiments
We conducted the experiments to verify the performance of proposed LSTM-based method. The experimental conditions were designed to confirm the effectiveness of pre-processing and processing components described in the methods section. As for pre-processing component, we verified that oversampling the data set in the pre-processing part could increase the performance of the informal abbreviation classifier. As for processing component, we verified that concatenating the word’s representation such as BoW and word2vec in the embedding layer of the LSTM model could increase the performance of the informal abbreviation classifier.
We evaluated the precision of the classifier when the abbreviation samples are added, in comparison to the basic model. We also evaluated the precision when the word representation input such as BoW and word2vec are added as a multiple features inputs to the model.
Data Set
Our data was acquired from a publicly available medical notes corpus [30] in English. The data consist of medical notes of varied types from different medical specialties, such as progress notes, SOAP notes, discharge summaries, etc [31]. This corpus is quite general, and contains informal abbreviations, including various writing styles, such as upper and lower case only and mixed cases, alphanumeric, and alphanumeric with symbolic characters.
Table 1 shows the distribution of abbreviation words and nonabbreviation words in the data set. A total of 475 sentences were extracted from randomly selected medical notes. A 475 sentences are able to represent most of the informal abbreviations for our case study. Using the preprocessing strategies we acquired 7159 samples; 704 of those were abbreviation samples (10%) and 6455 were non-abbreviation samples (90%). Furthermore, the vocabulary extracted from the data set consisted of 1629 unique words, including abbreviation words.
| Free text data | Size | Example |
|---|---|---|
| Sentences | 475 sentences | "coronary artery disease history of seizure disorder GERD bipolar" |
| Unique vocabularies | 1629 vocabs | PSA, GERD, mg, TEST, Cardiolite, cm, Dr, STRESS |
| Abbreviation words | 279 words | Q-fever, h/o, mg, G3P3, PSA, PMH, DVT, ml, cm, Dr |
| Non-abbreviation words | 8786 words | TEST, STRESS, incidental, DATA, Cardiolite |
| 5-word abbreviation samples | 704 samples | "of seizure disorder GERD bipolar" |
| 5-word non-abbreviation samples | 6455 samples | "coronary artery disease history of" |
Table 1: Data set distribution.
Each sample consists of a five sequence of words. We use the short sequence length, such as 5-words sample because in the classification task such as our case, it is easier to classify using shorter sequence length than longer sequence length [32]. The 7159 5-word samples were split into training data set (80%) and test data set (20%). Both data sets had the same proportion of abbreviation samples. This ratio could be specified as a parameter of the samples generator.
Model Implementation
We use bidirectional LSTM with forward and backward sizes 32, dropout of 0.3, and two dense layers, the first one with size 32 and ReLU [33] activation (to preserve the input’s sparsity), and the last one with activation softmax [34] for the final prediction with size 3. A batch size of 32 and 200 epochs were used for training.
From the same basic model, several combinations of inputs were implemented and evaluated. In the first test, only the word indices converted into embedding vectors with size 100 was used. For the remaining tests, we evaluated the combinations of: pretrained BoW only; pre- trained word2vec only; each of these vectors concatenated to the embedding vectors; a combination of BoW and word2vec; and all vectors together. The BoW matix had size 1629 (same as the size of the vocabulary), and the word2vec matrix had size of 100.
In the processing part, we only modified the input structures with embedding replacement and concatenation between embedding, BoW, and word2vec. The remaining LSTM layout such as sizes of layers, dropout, dense and activation layers are the same for all input combinations. The concatenation between embedding vectors and additional matrices uses LSTM with same configuration.
Evaluation Method
In To evaluate the performance of our classifier, we analyzed it from the perspective of both pre-processing and processing components in our proposed method. From the pre-processing perspective, we evaluate the model’s performance based on the abbreviation samples added to the pre-processing part of our methods. We hypothesize that more balanced sample increases the performance of classifier. From the model’s processing perspective, we evaluate the model’s performance based on the all concatenation combination of the input matrices from the embedding layer of LSTM, pretrained BoW matrix, and pretrained word2vec matrix. We hypothesize that concatenation of inputted matrices increase the classifier’s performance.
Since the original data set suffers from the class imbalance, data sampling evaluation is required to understand the effect of increasing the number of samples. The informal abbreviation is a special case of Out-of-Vocabulary (OOV) word, which requires more features to differentiate it from the ordinary words, therefore the evaluation of input combinations was performed to assess the effect of inputs combination to the model.
We focused on increasing the model’s precision, since the cost of false positives is high. Due to the number of negative (nonabbreviation) samples is very large, then the model has low precision. The low precision will cause many words classified as abbreviations or high false positives. When false positives are too high, the system is not reliable anymore as a detection system. Precision is more related to the positive class (abbreviation class) than to the negative class (non-abbreviation class), since it measures the probability of correct detection of the positive class.
Using small windowed size sample, such as 5-words sample, the recall and F1-score are not as high as the precision because the effect of number of words that are analyzed in the context of an abbreviation word. When the window size is large, recall increases because words farther away from the abbreviation word will be taken into account and more abbreviation words are more likely to be found. Precision will be higher when the window size is small [35].
Results
The basic model in Table 2, which is the baseline, had precision of 68.6%. The basic model in Table 2 used only embedding as input to the model. The original data set contains 10% of abbreviation samples, these samples represented only 2% of abbreviation words in total, and we believe that was the reason for the low performance of this model.
| Abbr samples | Precision (Avg) | (SD) | Recall (Avg) | (SD) | F1-score (Avg) | (SD) |
|---|---|---|---|---|---|---|
| 10 percent (baseline) | 68.60% | 14.1 | 38.20% | 0.3 | 48.60% | 4.1 |
| 20 percent | 70.60% | 15.1 | 46.10% | 2.4 | 55.00% | 4.6 |
| 30 percent | 77.10% | 7.9 | 42.70% | 5.8 | 54.50% | 3 |
| 40 percent | 80.40% | 6.1 | 55.10% | 1.7 | 65.20% | 0.8 |
| 50 percent | 83.20% | 4.2 | 48.30% | 3.3 | 57.30% | 3.3 |
| 60 percent | 85.00% | 5.7 | 49.50% | 2.7 | 62.40% | 2.1 |
| 70 percent | 85.10% | 5.9 | 49.50% | 2.8 | 62.40% | 1.2 |
| 80 percent | 91.10% | 8.6 | 45.50% | 4.9 | 60.30% | 3.8 |
| 90 percent | 91.40% | 1.7 | 48.70% | 6.2 | 63.30% | 5.2 |
Table 2: Improvement using samples generator.
Performance using Samples Generator
In Table 2, we measure the precision’s average and its standard deviation, recall’s average and its standard deviation, and F1- score’s average and its standard deviation from five trials of each abbreviation samples modelling.
We used a samples generator to increase the precision of our baseline model, by adding more abbreviation samples to the original data set. Table 2 shows the precision improvement as the ratio of abbreviation samples is increased. For recall and F1-score, the 40 percent of abbreviation samples give the highest results.
We progressively increased the number of abbreviation samples in 10% steps, until an intended precision was reached. The enhanced data set contained 90% of abbreviation samples, representing 18% of abbreviation words in total.
When the abbreviation samples are 90% of the total samples, the new data set raised the precision of the model to 91.4%, recall was 48.7%, and F1-score was 63.3%. The precision was steady increased until its highest precision at 90% of abbreviation samples, while the recall and F1-score were fluctuating increased. The highest recall was 55.1% and F1-score was 65.2% were at 40% of abbreviation samples. The new data set is then used as a baseline for the next experiment using additional inputted matrices.
Performance using additional matrices
In Table 3, we measure the precision’s average and its standard deviation, recall’s average and its standard deviation, and F1-score’s average and its standard deviation from five trials of each input combinations modelling. Table 3 shows the precision improvement when the embedding is replaced with the BoW matrix, concatenation of embedding with BoW matrix, concatenation of BoW matrix with word2vec matrix, and concatenation of embedding, BoW matrix, and word2vec matrix.
| Inputs | Precision (Avg) | (SD) | Recall (Avg) | (SD) | F1-score (Avg) | (SD) |
|---|---|---|---|---|---|---|
| Embedding (baseline) | 91.40% | 1.7 | 48.70% | 6.2 | 63.30% | 5.2 |
| Word2vec | 74.80% | 2.6 | 55.80% | 2.7 | 63.80% | 1.2 |
| BoW | 91.60% | 2.4 | 43.20% | 1.5 | 58.70% | 1.3 |
| Embedding+word2vec | 84.70% | 5.7 | 45.60% | 1.7 | 59.20% | 1.5 |
| BoW+word2vec | 92.00% | 4.6 | 53.10% | 3.4 | 67.20% | 2.6 |
| Embedding+BoW | 92.60% | 8 | 48.80% | 6.6 | 63.30% | 3.7 |
| Embedding+BoW+word2vec | 93.60% | 2.7 | 57.60% | 8 | 68.90% | 5.5 |
Table 3: Improvement using additional matrices.
The recall and F1-score of the model increased when the embedding is replaced with the word2vec matrix, concatenation of BoW matrix with word2vec matrix, and concatenation of embedding, BoW matrix, and word2vec matrix. For the concatenation of embedding with BoW matrix, the recall was slightly increased, while the F1-score was not changed.
The BoW matrix increased the precision of baseline model with the 90% of abbreviation samples to 91.6%. The BoW matrix and word2vec matrix concatenation increased the precision to 92.0%. The input addition of BoW matrix to the embedding of baseline model increased the precision to 92.6%. Finally, concatenation of embedding, BoW matrix, and word2vec matrix increased the precision to 93.6%. For recall, the word2vec matrix increased the model’s recall to 55.8%, the concatenation of BoW and word2vec increased the model’s recall to 53.1%, the concatenation of embedding and BoW increased slightly the model’s recall to 48.8%, and the highest recall was 57.6% that achieved by concatenation embedding, BoW matrix, and word2vec matrix. The F1- score was increased by word2vec to 63.8%, by concatenation of BoW and word2vec to 67.2%, and the highest F1-score by concatenation embedding, BoW matrix, and word2vec matrix.
DISCUSSION
The precision of the results is lower than the preliminary evaluation. However, considering that the rate of negative The results showed that the LSTM based model could accurately detect abbreviations made in diverse writing styles on free text medical notes. In addition, our model was able to predict abbreviations even with a small number of abbreviation words in the data set.
The LSTM model’s ability to predict the abbreviations relies on the fact that it learns from the context in which words appear in sequences [36]. Most medical notes are written following a certain narrative, which creates a relevant context in the sequence of words [37], therefore, although some words are informal abbreviations, the LSTM is able to recognize them by learning from surrounding words [38].
The precision, recall, and F1-score of model increases by adding more abbreviation samples and concatenating inputted matrices. In our case, we focus on the precision of classifier, because it is meaningful for information extraction tasks that often demanding for a high precision [39]. The recall and F1-score are not as high as the precision, and it is common for the information extraction tasks with small windowed size sample, such as 5-words sample [35].
From the pre-processing perspective, we discuss the effect of sample population in relation with the model’s precision increment. From the processing perspective, we discuss the effect of additional pre-trained features in elation with the model’s precision increment.
Effect of Sample Population
In our analysis, we observed that the LSTM baseline model’s precision can be increased by adding more abbreviation samples as shown in Table 2. Every 10% of abbreviation samples addition, the precision increases from 0.3 to 6.5 points. The lowest precision’s increment is from 80% to 90% of abbreviation samples, while the highest precision’s increment is from 20% to 30% of abbreviation samples. Usually, LSTM has a good performance in large data sets with little class imbalance [40]. By combining a sliding window and a samples generator, it was possible to increase the number of samples, while reducing the class imbalance, which allowed for an increase on the model’s precision.
Effect of Additional Pre-trained Features
It was also possible to increase the LSTM baseline model’s precision by including additional pre-trained features as a complementary input as shown in Table 3. The addition of BoW to the model’s input increases the precision from 0.2 to 2.2 points. The addition of word2vec increases the model’s precision if it also includes the BoW matrix addition. This takes advantage of the fact that the LSTM model can process multiple features for its learning [22]. Initially, the basic input-embedding vectors is used to differentiate between the classes in the binary classification (abbreviation vs nonabbreviation). As additional features are added to the LSTM model, it can improve its ability of differentiating between classes. For this end, word frequency (BoW) and word2vec vectors are commonly used features in natural language processing [41].
Adding the BoW increased the model’s precision. When using a pre-trained BoW instead of the word2vec, an even higher performance could be obtained. This result shows that the LSTM model for the detection of informal abbreviations is better at learning the information about word frequency represented by the BoW matrix, when compared to the information about word2vec matrix. This is due to the small number of abbreviation words in the data set, which makes the model separate clearly between abbreviation words that have low word’s frequency and non-abbreviations words, which have higher word’s frequency. In comparison with the word2vec, the small number of abbreviation words makes the model difficult to differentiate between abbreviation words and non- abbreviation words due to the less contexts available from the corpus data set.
Detection of informal abbreviation to achieve high degree of semantic interoperability
Interoperability standards, such as SNOMED CT, state that medical terms should be expressed in a structured and unambiguous way, providing a semantic expression whose meaning can be agreed upon by different systems, in a consistent and clearly expressed manner.
However, in the case of free text medical notes, medical expressions are sometimes written using informal abbreviations that may not be detected by SNOMED CT based processors. This could negatively impact patient care and generate burden for third party usage [42].
It is not sufficient that the physician who wrote the medical note is able to understand it, multiple people who participate in the medical care process must understand them and agree on the meaning of all terms. In this sense, if informal abbreviations, such as those shown in Table 1, can be clarified in a disambiguation process, higher degrees of semantic interoperability can be achieved.
This allows relevant clinical information to be recorded using consistent, common representations during a consultation and supports the sharing of appropriate information with others involved in delivering care to a patient.
Limitations
This study had the following limitations. First the data set was composed of random samples from a public medical notes database, which may not fully represent all the writing styles found on real world medical notes. To further validate and improve the results, it is necessary to use larger data sets with medical notes from different hospitals and clinical settings. Additionally, the data set itself was not so big in comparison to other studies on natural language processing. However, in the specific context of medical notes, this is expected, since, due to issues of privacy and confidentiality, most studies can only have access to a limited number of real life medical notes. In previous studies, mtsamples. com data set are utilized as a bench-marked of clinical study, such as for extracting family history diagnosis from clinical texts [43] and for identifying representation of drug use [44]. In addition, because one of our evaluation is to evaluate the classifier to detect informal abbreviations across many categories with class imbalance problem, such as the number of abbreviation words is far less than the number of non- abbreviation words, the public data set such as mtsamples.com fit properly for our study. We argue that if the classifier has high performance in detecting abbreviations with a far less population, then it should perform better for detecting a higher population of abbreviations that commonly found in the real data set.
Generality
This study can be extended to detect multiple undefined medical entities from free text medical notes, such as symbolic terms, for example “ ? ,, -->’’ (+)“,” ? ,, -->’’ (-)“, and “ ? ,, -->’’ (↑)“, with diverse variation of writing styles similar to the informal abbreviations cases. A symbolic terms are kind of abbreviated version of common words that are often used in the medical notes to express the patient’s condition, such as “ ? ,, -->’’ (+)“ for “ ? ,, -->’’positive“,” ? ,, -->’’ (-)“ for “ ? ,, -->’’negative“, and “ ? ,, -->’’ (↑)“ for „increased“.
CONCLUSION
Using the LSTM model is innovative to recognize informal abbreviations with diverse writing styles that are commonly found in the free text medical notes. Our approach differs with the previous approach that only focus on the formal abbreviation with the predefined features [9]. Using our approach, the precision for the informal abbreviation detection is 93.6%, recall is 57.6%, and F1- score is 68.9%, with using only small population of data set. Thus, our study will provide a case study for future NER researches using small data set. We use the precision as an evaluation metric because we want to minimize false positives or to optimize the specificity of our classifier. Overall, other metric such as recall and F1-score can be still be used if necessary.
For possible future extension of our work, it will be useful to develop a writing support systems, which can recognize informal abbreviations in real-time while the physician is typing it and raise appropriate indicators for the informal abbreviation meaning confirmation, thus increase the semantic interoperability for EMR system.
Acknowledgements
This study is supported by JICA Innovative Asia Scholarship and partially supported by JSPS KAKENHI grant number 19K12820.
REFERENCES
- Lehne M, Sass J, Essenwanger A, Schepers J, Thun S. Why digital medicine depends on interoperability. NPJ Digital Med 2019; 2(1):1-5.
- Reisman M. EHRs: The challenge of making electronic data usable and interoperable. Pharmacy and Therapeutics 2017; 42(9): 572.
- Rodrigues JM, Schulz S, Rector AL, Spackman KA, Üstün B, Chute CG, et al.. Sharing ontology between ICD 11 and SNOMED CT will enable seam- less re-use and semantic interoperability. InMedInfo 2013: 343-346.
- Ross MK, Wei W, Ohno-Machado L. “Big data” and the electronic health record. Yearbook Medical Informatics 2014; 23(1): 97-104.
- Sheppard JE, Weidner LC, Zakai S, Fountain-Polley S, Williams J. Ambiguous abbreviations: An audit of abbreviations in paediatric note keeping. Arch Dis Childhood 2008; 93(3): 204-206.
- Walsh KE, Gurwitz JH. Medical abbreviations: Writing little and communicating less. Arch Dis Childhood 2008; 93(10): 816-817.
- Ritter A, Clark S, Etzioni O. Named entity recognition in tweets: An experimental study. In Proceedings of the conference on empirical methods in natural language processing. Asso Comput Linguis 2011: 1524-1534.
- Moon S, McInnes B, Melton GB. Challenges and practical approaches with word sense disambiguation of acronyms and abbreviations in the clinical domain. Healthcar Inform Res. 2015 Jan 1; 21(1): 35-42.
- Wu Y, Rosenbloom ST, Denny JC, Miller RA, Mani S, Giuse DA, et al. Detecting abbreviations in discharge summaries using machine learning methods. In AMIA Annual Symposium Proceedings. American Medical Inform Asso. 2011: 1541.
- Yeganova L, Comeau DC, Wilbur WJ. Identifying abbreviation definitions machine learning with naturally labeled data. In 2010 Ninth International Conference on Machine Learning and Applications. IEEE 2010: 499-505.
- Katiyar A, Cardie C. Nested named entity recognition revisited. North American Chapter of the Association for Computational Linguistics: Human Language Technologies 2018; 1: 861-871.
- Yao L, Liu H, Liu Y, Li X, Anwar MW. Biomedical named entity recognition based on deep neutral network. Int J Hybrid Inf. Technol. 2015;8(8): 279-288.
- Yepes AJ. Word embeddings and recurrent neural networks based on Long-Short Term Memory nodes in supervised biomedical word sense disambiguation. J Biomed inform. 2017;73: 137-47.
- Li J, Sun A, Han J, Li C. A survey on deep learning for named entity recognition. IEEE Transactions on Knowledge and Data Eng 2020.
- Yang Z, Dehmer M, Yli-Harja O, Emmert-Streib F. Combining deep learning with token selection for patient phe- notyping from electronic health records. Scientific Reports 2020; 10(1): 1-8.
- Lyu C, Chen B, Ren Y, Ji D. Long short-term memory RNN for biomedical named entity recognition. BMC Bioinformatics 2017; 18(1): 462.
- Rondeau MA, Su Y. LSTM-Based NeuroCRFs for Named Entity Recognition. NTERSPEECH 2016: 665-669.
- Magge A, Sarker A, Nikfarjam A, Gonzalez-Hernandez G. Comment on: “Deep learning for pharmacovigilance: recurrent neural network architectures for labeling adverse drug reactions in Twitter posts”. J American Medical Inform Ass. 2019; 26(6): 577-579.
- Derczynski L, Maynard D, Rizzo G, Van Erp M, Gorrell G, Troncy R, et al. Analysis of named entity recognition and linking for tweets. Info Process Manage 2015; 51(2): 32-49.
- Han H, Wang WY, Mao BH. Borderline-SMOTE: A new over- sampling method in imbalanced data sets learning. In International conference on intelligent computing. Springer, Berlin, Heidelberg; 878-887.
- Sahare M, Gupta H. A review of multi-class classification for imbalanced data. Int J Advan Comp Res 2012; 2(3): 160.
- Li H, Shen Y, Zhu Y. Stock price prediction using attention- based multi-input LSTM. In Asian Conference on Machine Learning 2018: 454-469.
- Wang Q, Downey C, Wan L, Mansfield PA, Moreno IL. Speaker divarication with lstm. Conference on Acoustics, Speech and Signal Processing (ICASSP). IEEE. 2018: 5239-5243.
- Wang P, Qian Y, Soong FK, He L, Zhao H. A unified tagging solution: Bidirectional lstm recurrent neural network with word embedding. 2015.
- Graves A, Fernández S, Schmidhuber J. Bidirectional LSTM networks for improved phoneme classification and recognition. Int Conf Artificial Neural Networks. 2005: 799-804.
- Ruch P, Baud R, Geissbühler A. Evaluating and reducing the effect of data corruption when applying bag of words approaches to medical records. Int J Med Inform. 2002; 67(1-3): 75-83.
- Rong X. word2vec parameter learning explained. arXiv 2014.
- Pennington J, Socher R, Manning CD. Glove: Global vectors for word representation. InProceedings of the 2014 conference on empirical methods in natural language processing (EMNLP) 2014: 1532-1543.
- Sundermeyer M, Schlüter R, Ney H. LSTM neural networks for language modeling. Thirteenth annual conference of the International speech communication association 2012.
- Lewis N, Gruhl D, Yang H. Extracting Family History Diagnosis from Clinical Texts. InBICoB 2011: 128-133.
- Kobayashi S, Kume N, Nakahara T, Yoshihara H. Designing Clinical Concept Models for a Nationwide Electronic Health Records System For Japan. EJBI. 2018; 14 (1): 16-21.
- Bermingham A, Smeaton AF. Classifying sentiment in microblogs: Is brevity an advantage?. InProceedings of the 19th ACM International conference on Information and knowledge management 2010: 1833-1836.
- Agarap AF. Deep learning using rectified linear units (relu). arXiv 2018.
- Zhao D, Wang J, Lin H, Yang Z, Zhang Y. Extracting drug–drug interactions with hybrid bidirectional gated recur- rent unit and graph convolutional network. J Biomed Inform 2019; 99: 103295.
- Smith L, Tanabe LK, Ando RJ, Kuo CJ, Chung IF, Hsu CN, et al. Overview of BioCreative II gene mention recognition. Genome Bio. 2008; 9(2): S2.
- Bowman SR, Gauthier J, Rastogi A, Gupta R, Manning CD, Potts C. A fast unified model for parsing and sentence understanding. arXiv 2016.
- Leroy G, Chen H, Martinez JD. A shallow parser based on closed-class words to capture relations in biomedical text. J Biomed Inform 2003;36(3): 145-58.
- Baziotis C, Pelekis N, Datastories DC. Deep lstm with attention for message-level and topic-based sentiment analysis. In Proceedings of the 11th International workshop on semantic evaluation. 2017: 747-754.
- Klinger R, Friedrich CM. User’s choice of precision and recall in named entity recognition. InProceedings of the International Conference 2009: 192-196.
- Sanabila HR, Jatmiko W. Ensemble learning on large scale financial imbalanced data. In2018 International Workshop on Big Data and Information Security (IWBIS). IEEE 2018: 93-98.
- Turner CA, Jacobs AD, Marques CK, Oates JC, Kamen DL, Anderson PE, et al. Word2Vec inversion and traditional text classifiers for phenotyping lupus. BMC Med Inform Decision Making. 2017; 17(1): 126.
- Liu H, Wu ST, Li D, Jonnalagadda S, Sohn S, Wagholikar K, et al. Towards a semantic lexicon for clinical natural language processing. InAMIA Annual Symposium Proceedings. J American Medical Inform Ass. 2012: 568.
- Bill R, Pakhomov S, Chen ES, Winden TJ, Carter EW, Melton GB. Automated extraction of family history information from clinical notes. InAMIA Annual Symposium Proceedings. J American Medical Inform Ass 2014: 1709.
- Carter EW, Sarkar IN, Melton GB, Chen ES. Representation of drug use in biomedical standards, clinical text, and research measures. InAMIA Annual Symposium Proceedings. J American Medical Inform Ass 2015: 376.